Wikidata in Stardog

Wikidata is a free and open knowledge base that can be read and edited by anyone. It serves as the central storage for the structured data of Wikipedia and other Wikimedia projects like Wiktionary, Wikisource, and more. It is also one of the largest publicly available RDF datasets and exports of the complete dataset are provided daily. As of this writing in January 2022, the Wikdiata RDF export contains 16.7 billion triples for close to 100 million entities and the size of the dataset continuously grows.
Wikidata provides an online query service where anyone can write a SPARQL query. Not surprisingly, the data model is large and complex but there are many example queries provided on the Wikidata website to get you started.
Wikidata query service naturally defines certain time and resource restrictions on the queries you can run. If you would like to run more complex queries or use Wikidata in your application in a reliable way you would prefer to have the data loaded locally and avoid the risks associated with an external service.
In this post, we will describe our experiments for loading Wikidata into Stardog and querying it so you will now what to expect. Here is a TLDR of the results:
- Stardog can load the complete Wikidata dataset (16.7 billion triples) under 10 hours at a rate of roughly half a million triples per second on a commodity server
- The average query execution time for the queries in the Wikidata Graph Pattern Benchmark (WGPB) against Wikidata is under 100ms and 99% of the queries finish under 1 second
In the rest of this post we will explain the details of our experiments so you can reproduce our results. Let’s get started.
Loading Wikidata
As we talked about in the past, Stardog is highly optimized for loading large datasets. Similar to previous experiments, we followed the general performance tuning suggestions we describe in our docs. There are couple minor tweaks specific to Wikidata as explained below.
First, for hardware, we went with a memory-optimized EC2 instance but opted to use less expensive (and less performant) SSD disks:
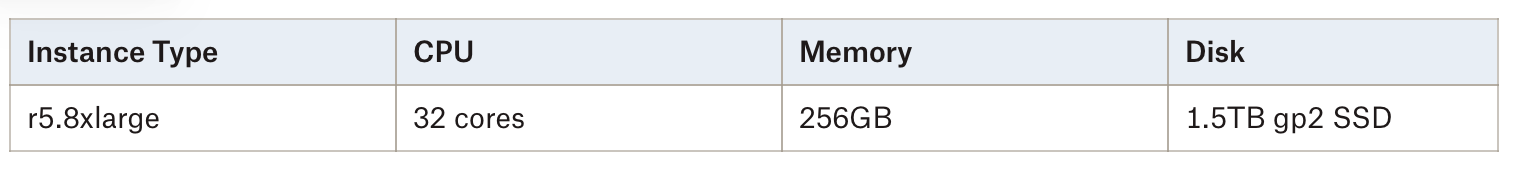
The performance numbers we present here would improve with compute-optimized instances and/or higher throughput io1 SSDs but we wanted to show that you can get very good results without going to higher-end hardware.
We used the memory settings -Xms30g -Xmx30g -XX:MaxDirectMemorySize=200g for the JVM and set memory.mode=bulk_load in stardog.properties to improve loading performance. We used the db create command with the following options:
stardog-admin db create -o strict.parsing=false index.statistics.chains.enabled=false -n wikidata <input-path>
We had to disable strict parsing because Wikidata includes some dates that are not valid. By default Stardog would refuse to load such data but when strict parsing is disabled these dates would be loaded into the database and can be detected and fixed later. Chain statistics is a summary of most frequent chains occurring in the data. It is used by the query engine to optimize complex SPARQL queries. Frequent chain mining sometimes takes a long time on very large and complex graphs but it is not required for most queries and can be done asynchronously if necessary.
As we discuss in our docs, loading performance will be maximized when the input data is partitioned into multiple files so multiple cores can be used for parsing the input files. When you have control over how the data is generated you can easily generate multiple files. However, in the case of Wikidata, the RDF data is provided as a single file so splitting should be done as an additional step.
We downloaded the latest-all.nt.gz file (178GB) and used the following command to split it into smaller files with 150M triples in each:
$ gunzip -c latest-all.nt.gz | split -l 150000000 -a 3 -d --additional-suffix .nt --filter='gzip > $gz'- wikidata-
Note that the --filter argument ensures the split output will be compressed and won’t use too much disk space (uncompressed Wikidata RDF file takes 2TB disk space). Note that using NTriples syntax ensures split files are valid too. If you download the Wikidata in Turtle format you cannot split it into multiple files as easily because output would be syntactically invalid.
Splitting the input file speeds up loading but this additional preprocessing step itself takes 15 hours. In the upcoming 7.9.0 version we are introducing a new optimization that can parse a single file in multiple threads to eliminate the additional preprocessing step. This optimization works both for the NTriples format and the NQuads format.
The following chart shows the Wikidata loading times for the Stardog 7.8 version and the upcoming Stardog 7.9 version both when the Wikidata is partitioned into multiple files before loading and when it is loaded as a single input file.
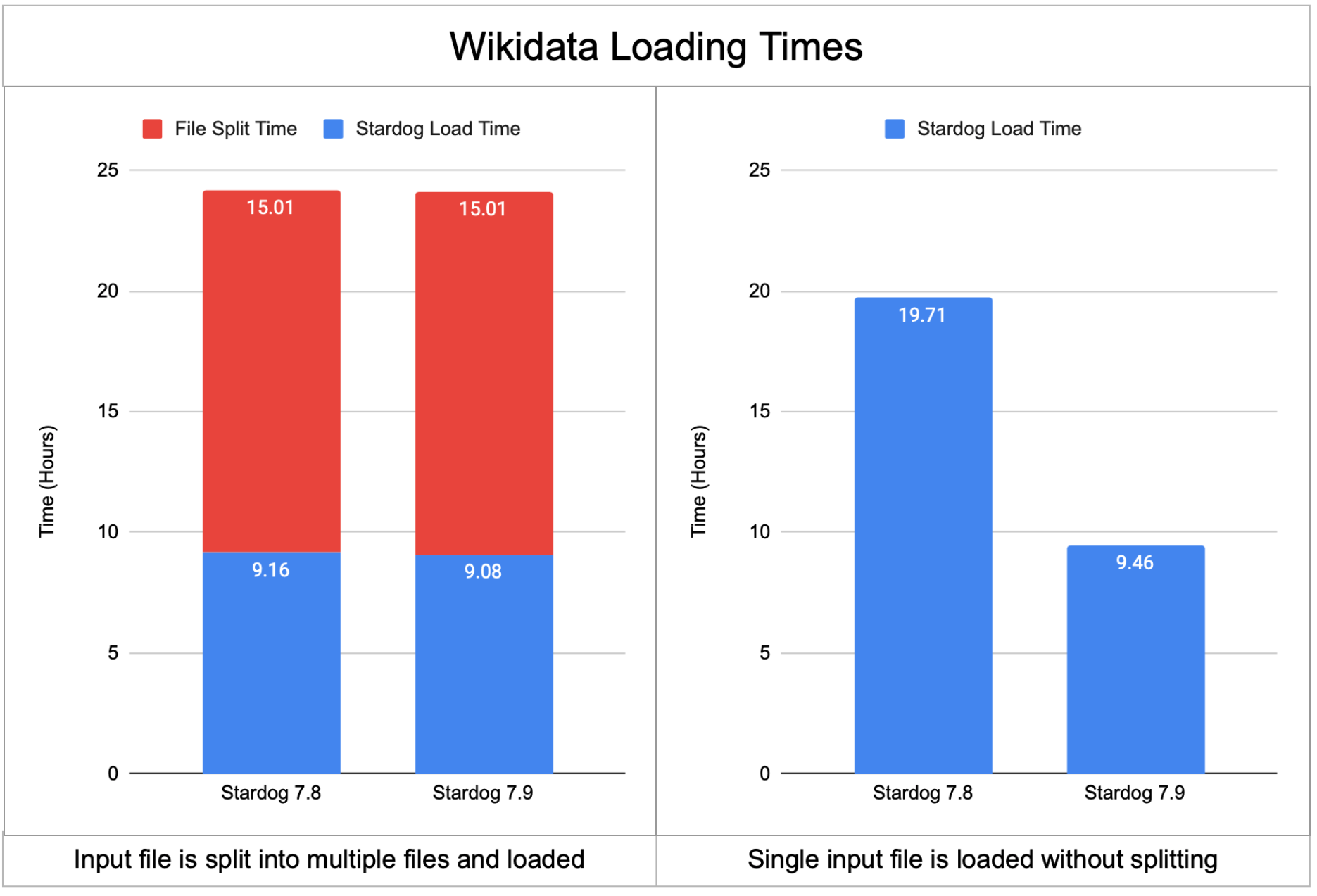
The loading time did not change between 7.8 and 7.9 when input is in multiple files but when the input file is provided as a single file loading time is decreased by half. The more detailed timings are provided in the following table:
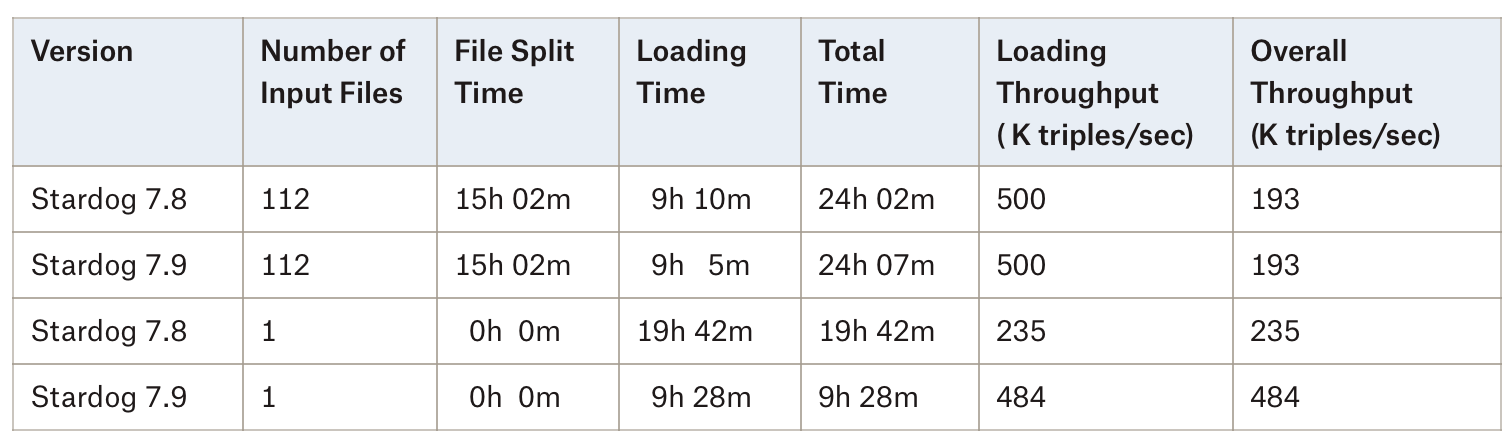
As these results show, loading is still slightly faster when the input is provided as multiple files but with 7.9 there is not a big penalty anymore if the input is not partitioned and you can save a lot of time by not splitting files.
Querying Wikidata
Once the data is loaded you need to remove the memory.mode=bulk_load setting from stardog.properties and restart the server. You can then start running queries but beware that for such large databases first few queries will be slow until the storage caches are warmed up. Here is an example query provided by Wikidata:
#People that received both Academy Award and Nobel Prize
SELECT DISTINCT ?Person ?PersonLabel ?NobelPrizeLabel ?AcademyAwardLabel WHERE {
?NobelPrize wdt:P279?/wdt:P31? wd:Q7191 . # <- subtypes of nobel prize
?AcademyAward wdt:P279?/wdt:P31? wd:Q19020 . # <- subtypes of academy award
?Person wdt:P166 ?NobelPrize . # <- people awarded a nobel prize
?Person wdt:P166 ?AcademyAward . # <- people awarded an academy award
?Person rdfs:label ?PersonLabel FILTER(lang(?PersonLabel) = "en")
?NobelPrize rdfs:label ?NobelPrizeLabel FILTER(lang(?NobelPrizeLabel) = "en")
?AcademyAward rdfs:label ?AcademyAwardLabel FILTER(lang(?AcademyAwardLabel) = "en") }'
You will notice that the original version of this query as well as other examples provided by Wikidata includes a non-standard SPARQL service denoted as SERVICE wikibase:label. The purpose of this service is to automatically return the labels associated with entities in the result set. In our version we added extra patterns to the query to retrieve the labels in English. Obviously the filter can be modified to select another language as Wikidata contains labels in many languages.
We have tried several of the example queries and have seen pretty good query performance especially when compared with the online query service. But we wanted to go beyond anecdotal evidence and run more complete benchmarks as we explain next.
Benchmarking Wikidata
One benchmark we have identified is the Wikidata Graph Pattern Benchmark (WGPB) for RDF/SPARQL developed by researchers at the University of Chile. This benchmark was developed Aidan Hogan et al. for their paper “A Worst-Case Optimal Join Algorithm for SPARQL". The goal of the benchmark is to test the performance of query engines for more complex basic graph patterns. It includes 50 instances of 17 different query patterns resulting in a total of 850 queries. The query patterns range from joins with a single variable to triangle patterns to square patterns.
The WGPB benchmark was used to evaluate different SPARQL engines on a much smaller subset of the Wikidata graph with roughly 90M triples but we went ahead and tested these queries against the complete 16.7B triples which is 185 times bigger! During our initial experiments we observed that Stardog’s query plan cache feature was not working very well with the default settings because there were many queries with structurally similar patterns yet widely different selectivity. We have set the database option to query.plan.reuse=CARDINALITY which caused cardinality estimations to be taken into account before reusing query plans which resolved the problem.
For our experiments we have run 850 queries 10 times. For each iteration, we randomly shuffled the queries so there wasn’t a regular pattern to how queries were executed. No query results were cached between iterations, only the query plans were reused by the engine when needed. The results for the first 2 iterations were excluded as warmup runs and we computed the average execution times for each query.
There are too many queries in the benchmark to present the execution times individually so we summarize the results in the following histogram:
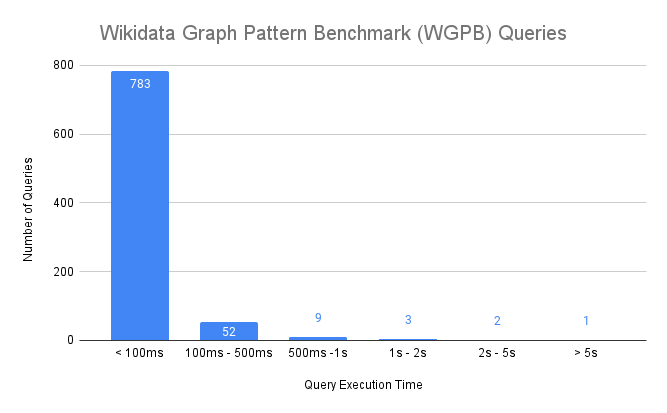
Here are some key takeaways:
- 783 queries (92% of all queries) finish under 100 milliseconds
- 844 queries (99% of all queries) finish under 1 second
- Only 1 query takes more than 5 seconds to complete
- The relative standard deviation for query execution times (not shown above) was less than 10%
Overall these query results are pretty impressive even when compared to the original WGPB experiments that were run against a much smaller version of the Wikidata graph.
Try it yourself!
The best thing about benchmarking with a public dataset is you can download the data and verify the results yourself. If Wikidata is of interest to you give it a try and let us know how it went in the Stardog Community.
Keep Reading:

Unified Process Monitoring
Did you ever have a slow query, export or any other operation and wondered “Is this ever going to finish?!”. Many operations such as SPARQL queries, db backups etc have performance characteristics which may be hard to predict. Canceling rogue queries was always possible, but there was not yet a way to do so for other potentially expensive operations. If this ever affected you then the new unified process monitoring feature we added in Stardog 8.

Chaos Testing Stardog Cluster for Fun and Profit
At Stardog we work hard to build software that’s not only performant but also extremely robust. Stardog Cluster is a highly available, key component to ensuring Stardog remains up and running for our customers. However, every distributed system is susceptible to faults and bugs wreaking havoc on it. Most software testing checks the “happy path” through the code to verify it behaves as designed. Moving a step further, tests can check some obvious failure cases; for example, if one service in a deployment is terminated.
Try Stardog Free
Stardog is available for free for your academic and research projects! Get started today.
Download now