Today, researchers have access to more data than ever; including genomics, clinical trial, social media, wearables, ingestibles, and other real world data. This sea of data, filled with promise, also introduces new problems.
The content and structure of this data varies widely. Because researchers cannot easily integrate this data, they are left to process data manually or rely on tribal knowledge. This prevents researchers from unlocking the potential of their data to accelerate drug discovery. Scoping new experiments requires manual research and data preparation to gather relevant previous studies and experiments. As a result, researchers are slow to form hypotheses for the basis of experiments; wasting time and resources on inefficient or ill-formed experiments.
The goal: quickly ask complex questions
Instead, what if R&D groups could quickly ask complex questions?
Show me all the studies where subjects/groups were treated first with dosed compound 1 and subsequently by compound 2.
What have we done on osteoarthritis in the last 10 years? At which sites?
Were there patients in clinical trials with a profile similar to a field safety report?
Which set of compounds are creating a similar effect or which compound has been tested in similar conditions and similar treatments?
Could some gene expression be used as a biomarker to understand whether some drug is delivering some effect?
It’s possible to uncover the relationships amongst all R&D data to be able to answer these questions and more. However, the complexity of the data itself has often hindered digital transformation efforts.
What makes this data so hard to integrate?
It’s not uniform. This data is highly varied — IoT data, social media posts, doctor’s notes, clinical trial results, and more. This data is ultimately a mix of structured, semi-structured, and unstructured data, and most data integration platforms are not designed to accommodate data from such a diversity of sources.

Relationships are complex. The interplay between genes, targets, adverse effects, diseases, symptoms, trials, sites, and more is incredibly complex. Not to mention added complexity once a drug goes into production. A molecule may be renamed in trials, branded upon drug release, and even rebranded later on. How then can you tie a complaint about a certain drug back to the manufacturing site of a particular drug ingredient? A holistic data solution for pharma must be able to address this complexity with precision.
It’s not consistently named. Any given gene will have different ID numbers in different labs. Multiply this by all the entities required for R&D — genes, expressions, adverse effects, and more. This becomes a monumental problem at the scale of global pharmas, who may be evaluating data from dozens or hundreds of labs. It’s impossible to enforce terminology consistency across each lab, which operates with some level of independence, not to mention external labs and datasets.
It’s physically distributed. Data within an organization is stored within many different systems. A system that manages site enrollment is separate from the system used for study design. Yet another system is used for electronic data capture, and so on. When companies need to bring data together to solve a specific problem, they often bring it together in yet another relational system. This simply leads to more copies of the same data, which introduces an accompanying increasing risk of human error.
It’s hard to navigate. Unfortunately, even if you bring all your data together, it doesn’t solve the problem. Data lakes alone have not led to better clinical outcomes. Data scientists are still mired in exploring and preparing data for each new analysis.
Plus, it’s not just internal data that matters. External data, from clinical trial studies to reference data, is critical to R&D workflows, from leveraging academic collaborations to informing chemists what avenues have been pursued by other companies to enriching competitive intelligence. If researchers are looking for all trials involving a particular molecule, it can be difficult to know they’ve consulted all relevant research. For better patient outcomes, the data must be simpler to navigate.
The future of R&D requires connected data
To solve these problems, R&D operations must focus on a flexible, reusable data foundation. The solution must involve data that can accurately reflect the complexity of real-world operations, down to computational biology and precision medicine.
Connected data, enriched with meaning, is the key to the new digital foundation of R&D. Connected data is the output of knowledge graphs, which support integration of varied data types and flexible data models that allow for multiple interpretations of the same data. If you are curious about how knowledge graphs work, see, “What is a Knowledge Graph?”
Connected data represents the real world. The real world doesn’t operate within the confines of tables. Connected data easily represents complex relationships, accurately reflecting the drug development process and resulting data very precisely. Connected data also persists the metadata right alongside the data itself, preserving experiment conditions.
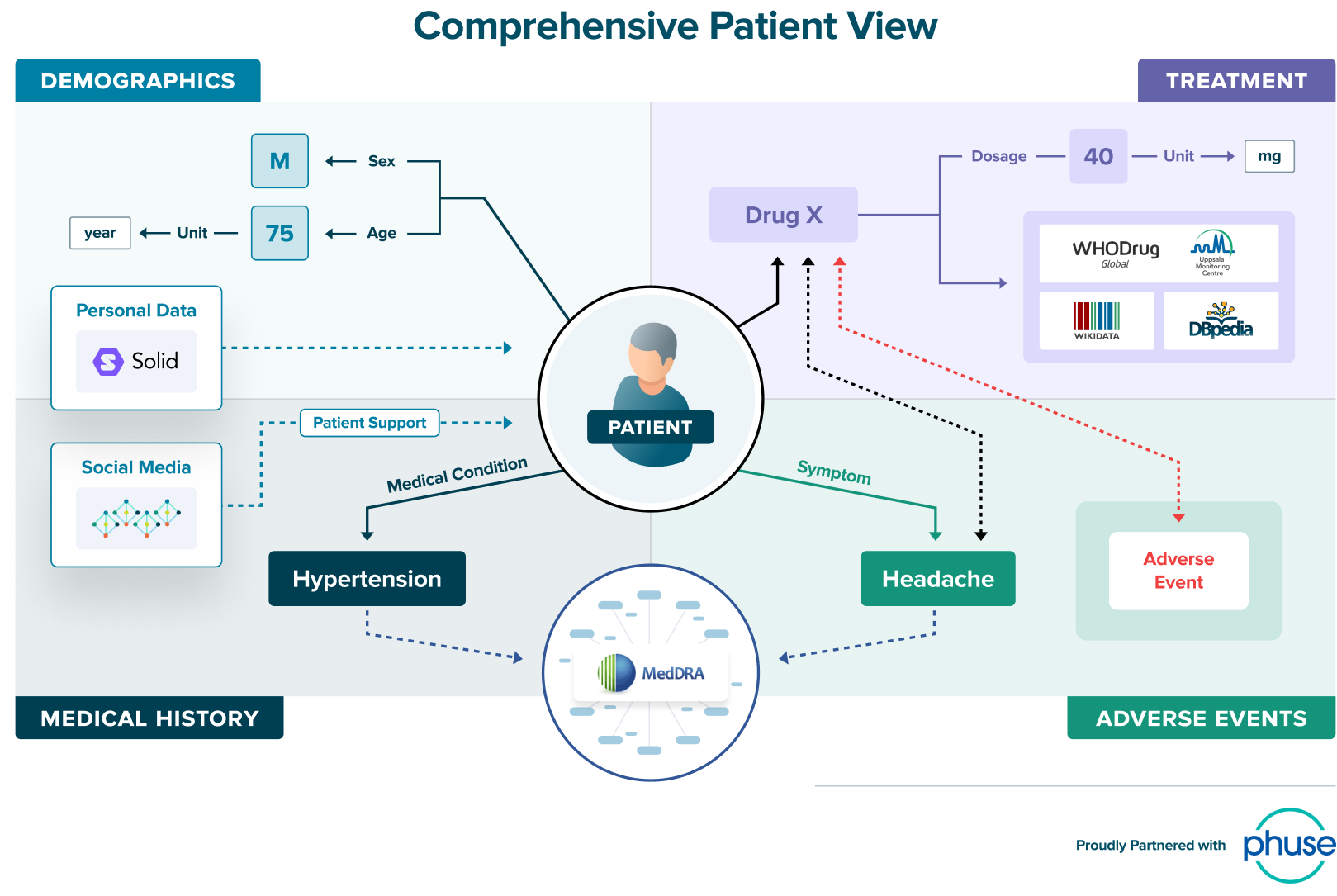
Slide source: Tim Williams, Phuse, adapted from webinar “How Knowledge Graphs will Transform the Pharmaceutical Industry”
Connected data is patient centric. Precision medicine is just one of the drivers towards patient centricity, alongside consumer demand and regulatory mandates. Inherently, connected data lends itself to patient centricity because it allows you to put the patient at the center of every analysis.
“If you have this mutation, you should be getting this treatment. If you have this other mutation, you should be getting a different treatment. Stardog allows us to tailor individual treatments, driving precision-medicine efforts forward.”
- Conrad Leonard, Senior Bioinformatician, QIMR Berghofer
Ultimately, the ability to represent the real world means that pharmaceutical organizations can be more patient centric. By creating a network of connected knowledge, all associations between various entities are captured. Learn more about how this unlocks capabilities for precision medicine for cancer genomics research group QIMR Berghofer.
Connected data provides traceability. Traceability is critical to empower data scientists and researchers to follow their instincts to understand the data. With Stardog you can follow the trail from the sample, to the sequencing library that was made from that sample, to the sequence that was made on that library, to data that came off the sequencing machine, to the next layer of analysis, and the next.
Connected data supports collaboration. Academic partners, external databases, licensing partners — collaboration of all forms is on the rise.
“Accenture research predicts New Science therapies will represent 54% of sales between 2017 and 2022, up from 47% between 2012 and 2017. Leaders in this space are investing heavily in emerging technologies to improve clinical outcomes and the patient experience, and connecting with an ecosystem of partners to accelerate innovation.”
At one global pharma, Stardog is used to support the analytics needs of the translational sciences and computational biology groups. 30% of active ingredients under evaluation were sourced from external collaborations, and there was limited control over the quality of this data. Further, there was a desire to incorporate publicly available gene expression research. Connected data fulfilled the need for a flexible solution that could relate their internal experiments to external and publicly available studies.
Connected data in practice
A top global pharmaceutical has dozens of labs across multiple continents. M&A activity further complicated their global footprint of labs. These labs all maintain their own research data, including information on compounds, tests, machinery, findings, and experiment conditions. But there was no repeatable way to connect the data from the labs together or connect the lab data with the internal and external reference data. This lack of connected data is compounded by the lack of consistent naming structures across the company. Different labs have different names or codes for the same compounds and so to combine the data, any given analyst had to know all the permutations of the same name or code.
That meant it was either impossible or incredibly time consuming for analysts on the business side to find the answer to their questions that span across labs — How much money are we spending to research a compound? How many tests have we run in the last 30 days? Further, they realized that business questions are always a chicken and the egg problem; as soon as you unify data to answer one question, there will be follow-up questions that will require data that hasn’t been unified yet. A responsive, agile solution was required.
Deploying Stardog to connect their lab data allowed all data to be unified virtually, leaving research data in place. Stardog linked related data, unifying all data and metadata without transformation. This allows analysts to easily see all the relevant data on a given compound, regardless of how it is referred to in individual labs or external sources. For business analysts, they created a Google-like semantic search interface for easy navigation. Now, business users and analysts can quickly answer cross-laboratory questions and create required reports.
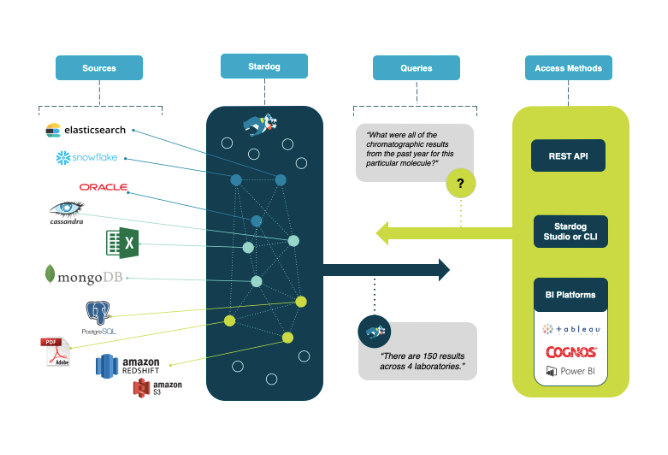
Stardog unifies data based on meaning, without requiring copying data. Ask complex questions in quick question-and-answer cycles to support research and business needs.
Trusted by global pharmaceuticals
Stardog’s Enterprise Knowledge Graph platform is uniquely suited to connect data and fulfill the needs of pharmaceutical organizations. Unifying relevant R&D data leads to faster drug target identification, and with metadata intact, researchers have the information they need to replicate experiments.
Stardog is in use by multiple top global pharmas, including Boehringer Ingelheim, Bayer, and many more. Interested to learn more? See how Stardog fulfills the promise of FAIR for longterm R&D success. Or, contact us to discuss ways to accelerate your drug discovery practices with Stardog.

