Make your Data FAIR

Get the latest in your inbox

Get the latest in your inbox
FAIR data principles serve as a north star for pharmaceutical R&D, promising accelerated innovation through findable, accessible, interoperable, and reusable data. Read on to learn how knowledge graphs make data FAIR, how knowledge graphs compare to other data management technologies, and how one leading global pharma succeeded in their FAIR transformation.
While FAIR data principles are well-established, connecting FAIR to business outcomes is critical to generate buy-in throughout an organization.
Here’s how FAIR can help:
The key is that FAIR data resolves the data complexities that result in lethargic and fractured efforts. With FAIR, you can create a digital foundation to support countless use cases alongside your ecosystem of collaborators.
A leading global pharmaceutical company realized they needed to shift from a reactive to a responsive data strategy to support the company’s R&D goals. FAIR was critical to supporting bioinformaticians within the translational sciences and computational biology divisions. While they had a data lake in place, their researchers were wasting time searching for the data they needed. The data they did have wasn’t being leveraged to its fullest potential. The goal was to provide data at a higher quality so that data scientists could focus on analysis, not data engineering.
A key consideration for their FAIR solution was that 30% of active ingredients under evaluation were sourced from external collaborations, and there was limited control over the quality of this data. Further, there was a desire to incorporate publicly available gene expression research. These requirements demanded a flexible solution that could relate their internal experiments to external and publicly available studies.
By implementing Stardog atop their data lake, they are creating a company-wide one-stop shop for 90% of their R&D data. Metadata on samples, studies, targets, labs and more is captured by Stardog, allowing researchers to ask questions such as:
Ontologies and vocabularies are used to identify synonymous genes and targets to make the dataset fully interoperable. Stardog provides high quality data access directly to data scientists, accelerating drug target identification and drug repurposing efforts.
“For us it was a natural choice to deviate from the pure data lake technologies to a more sophisticated model.”
- Head of IT Research Computational Biology and Translational Science
While many technologies from data catalogs to data lakes to data virtualization platforms promise FAIR, they don’t necessarily live up to the full mandate of FAIR. As a recap, here are the key principles of FAIR:
Findable: Metadata and data should be easy to find for both humans and computers. Machine-readable metadata are essential for automatic discovery of datasets and services.
Accessible: Once the user finds the required data, she/he needs to know how can they be accessed, possibly including authentication and authorization.
Interoperable: Data needs to be integrated with other data. The data must be interoperable with applications or workflows for analysis, storage, and processing.
Reusable: The ultimate goal of FAIR is to optimize the reuse of data. To achieve this, metadata and data should be well-described so that they can be replicated and/or combined in different settings.
Source: https://www.go-fair.org/fair-principles/
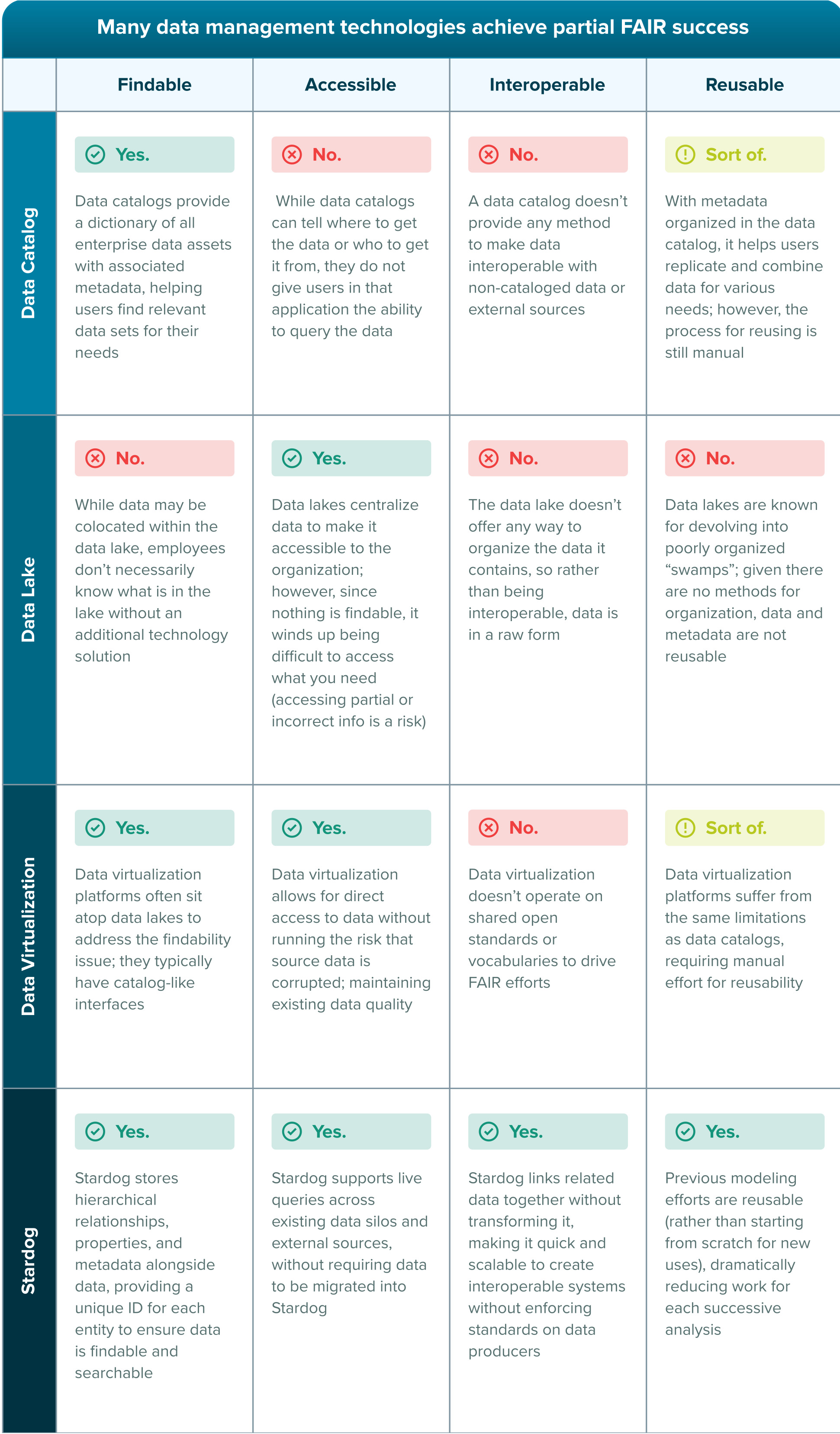

Knowledge graphs are consistently highlighted for their ability to deliver on FAIR data principles. While other data management technologies solve pieces of the puzzle, they are incomplete. Stardog’s Enterprise Knowledge Graph platform fully embraces the data management principles that FAIR promotes. But what exactly makes knowledge graphs the ideal FAIR solution?
FINDABLE
Many organizations are well on their way to having findable data thanks to data quality or cataloging initiatives. Stardog can access source systems using virtualization, connecting existing data source to the knowledge graph, without making a new copy of the data. Once data is connected to Stardog, you can map and model the data to link related data from various source data sets.
Stardog makes data findable whether you are starting with a data catalog, a data lake, or simply starting from scratch with various data silos. Regardless of your starting point, the first key to making data findable is representing the complex relationships that are rampant within pharma R&D.
Take a look at this comprehensive patient view which is enriched with data from both internal and external sources, and comes from data of varied structures, from databases to social media. This is what a knowledge graph looks like: a web of relationships.
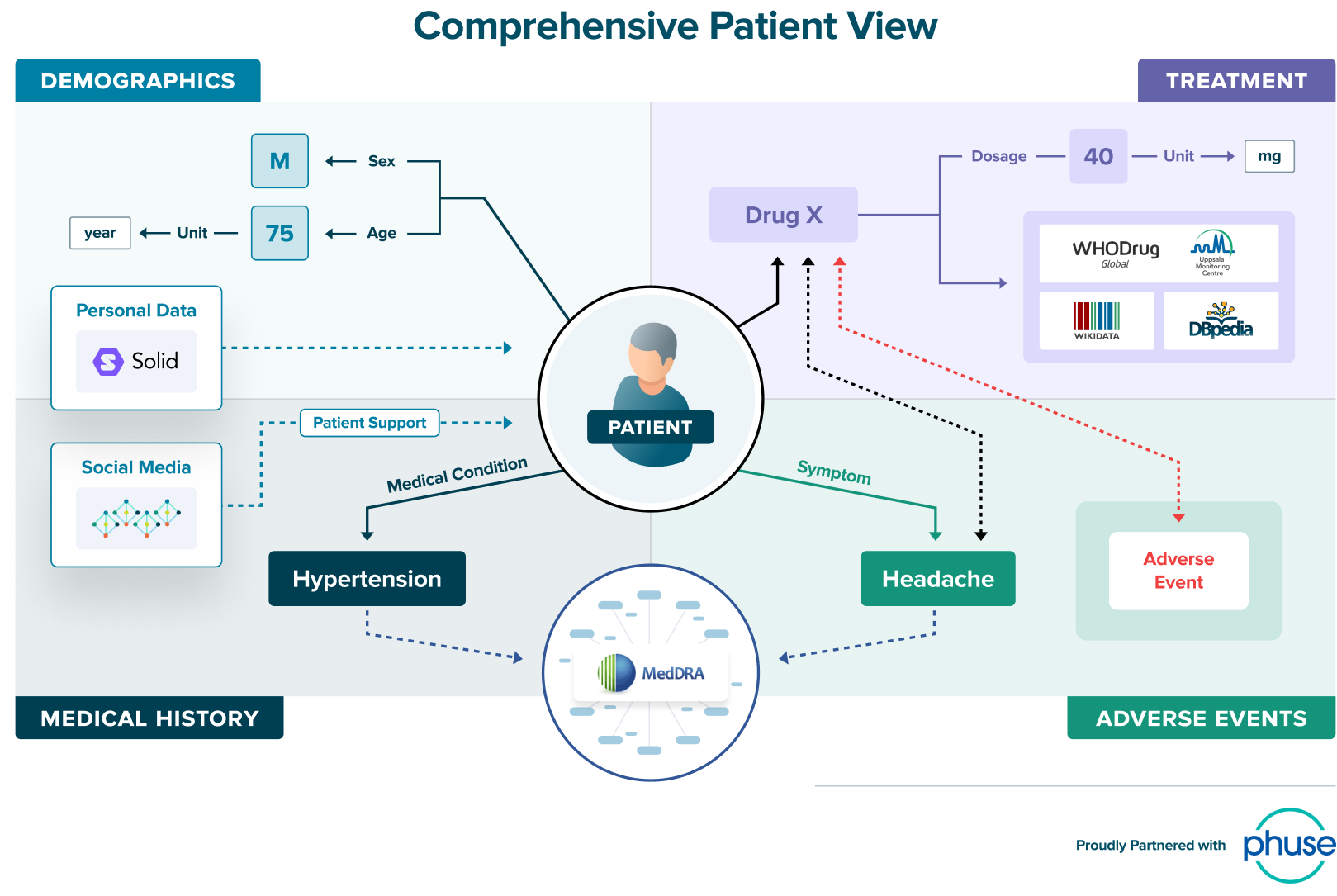
Slide source: Tim Williams, Phuse, adapted from webinar “How Knowledge Graphs will Transform the Pharmaceutical Industry.”
The second step to fulfilling findability is to comprehensively track metadata so that experiments can be easily reproduced. In Stardog, the metadata is stored alongside the data. The patient in the example above has a globally unique and persistent identifier, linking all related patient records to that ID. Since the metadata is stored alongside the data, both humans and machines can easily find relevant information required for analyses. This expressiveness is unique to knowledge graphs; any system based on relational data management would quite simply not be up to the task.
ACCESSIBLE
Data catalogs may help you understand the data you have, where it lives, and the status of data quality initiatives. However, most data scientists, researchers, or analysts want to access the data by asking questions and receiving answers. Unfortunately, data catalogs don’t make the data actually accessible within the platform by any form of analytics or query capabilities.
In contrast, Stardog allows users to access data directly via queries across data silos. Compared to relational systems like most data virtualization platforms, Stardog’s graph-based system allows for complex questions evaluating many-to-many relationships. Stardog supports standard query languages including SPARQL, GraphQL, and Gremlin. Unified data can also be accessed via SQL, increasing the reach of your project to various internal audiences. Finally, the connected data is fully accessible via REST API to assorted application and analytics uses.
Because Stardog has the most advanced graph-based virtualization on the market, you can query live data silos without worrying about the freshness of the data. Access data from various internal sources as well as external parties in one comprehensive platform.
INTEROPERABLE
Stardog is specifically designed to support interoperability between data silos. The key to understanding how Stardog integrates data is to know that it links or connects related data, rather than transforming it. This method of linking data is unique to the semantic graph standards Stardog supports (and our Founders helped to develop!).
Each data object is assigned a unique ID, to which all related information is linked. This unique process allows data owners to maintain control of source data. For pharmaceutical companies, this is critical to support an ecosystem of collaborators who may refer to molecules, drugs, or compounds by different names. These differences are resolved easily via the data model, without forcing collaborators to change their operations or creating complex ETL workflows to prepare incoming data.
Stardog also supports common vocabularies that meet FAIR standards. Because Stardog supports semantic graph standards, you can easily leverage the work of other researchers or organizations. Stardog’s best-in-class Inference Engine has the most advanced capabilities on the market for processing complex ontologies such as SNOMED.
REUSABLE
A key benefit of knowledge graphs over other approaches is the richness of the metadata that can be captured. All relevant metadata is captured within the knowledge graph. Because data doesn’t need to be copied to be analyzed, organizations reduce the risk of data quality degradation.
To manage accuracy of data and metadata, Stardog offers constraints. Constraints can find inconsistencies across your data silos, flag conflicting data, or prevent the knowledge graph from accessing bad data. Constraints also support measuring the quality of the data, performing verification after an integration, and assisting in planning future improvement measures.
While FAIR standards discuss reusability purely in terms of metadata; at Stardog, we also think about reusability as it pertains to data modeling. There’s always more data — new external data streams, new data sources needed for the next release, new acquisitions with their own messes of data. Stardog’s extensible data model easily incorporates new sources while maintaining original schemas and metadata. Additionally, Stardog offers unique functionality to allow multiple data models to act upon the connected data. Stardog’s Inference Engine interprets business rules, data, and new facts at query time across multiple data models. This allows different parts of the enterprise to have different, even incompatible, views or lenses onto the common, underlying connected data.
Read more in our blog, “How Data Reusability Accelerates your Enterprise.”
Want to learn more? Watch our joint webinar with Tim Williams of PHUSE where we discuss how knowledge graphs are transforming the pharmaceutical industry. The webinar provides an introduction to knowledge graphs and covers how knowledge graphs support FAIR, patient centricity, and data quality efforts.
PHUSE is an independent, not-for-profit organization and platform for the discussion of topics encompassing the work of data managers, biostatisticians, statistical programmers and eClinical IT professionals. Learn more on the PHUSE website.
How to Overcome a Major Enterprise Liability and Unleash Massive Potential
Download for free